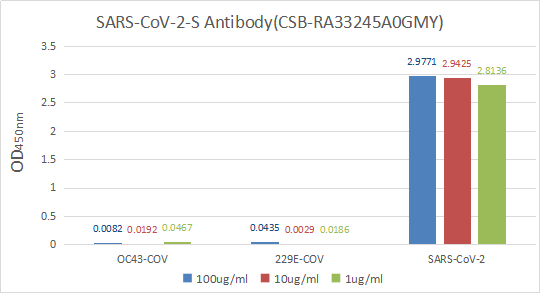
-
中文名称:S重组抗体
-
货号:CSB-RA33245A0GMY
-
规格:¥3080
-
图片:
-
其他:
产品详情
-
产品描述:CSB-RA33245A0GMY S重组单克隆抗体是针对S蛋白开发的高特异性科研试剂,采用重组技术确保批次间的高度一致性。该抗体靶向的S蛋白作为多种病原体表面的重要功能蛋白,在病原体入侵宿主细胞过程中发挥关键作用,是相关研究领域的重要检测靶点。经严格验证,本品在ELISA实验中展现出优异性能,推荐工作稀释度为1:10000至1:50000,可实现对目标蛋白的高灵敏度检测;在胶体金免疫层析(GICA)应用中同样表现卓越,推荐使用稀释梯度为1:500至1:25000,适用于快速检测体系的构建。其宽泛的有效稀释范围为实验设计提供了灵活选择,特别适用于体外检测、病原体相关蛋白的定性/定量分析、免疫层析试纸条开发等科研场景。产品经过多批次质控验证,确保不同实验条件下的稳定性和可重复性,为病毒感染机制研究、疫苗开发评估及快速检测技术研发等基础研究领域提供可靠的工具支持。
-
Uniprot No.:
-
别名:S; 2; Spike glycoprotein; S glycoprotein; E2; Peplomer protein)
-
反应种属:Human Novel Coronavirus (SARS-CoV-2/ 2019-nCoV)
-
免疫原:Recombinant Human Novel Coronavirus Spike glycoprotein (S) (16-685aa) (CSB-MP3324GMY)
-
免疫原种属:Human Novel Coronavirus (SARS-CoV-2/ 2019-nCoV)
-
标记方式:Non-conjugated
-
克隆类型:Monoclonal
-
抗体亚型:Monoclonal mouse (varialbe region) / human (kappa / IgG1 constant)chimeric antibody
-
纯化方式:Affinity-chromatography
-
克隆号:H6
-
浓度:It differs from different batches. Please contact us to confirm it.
-
保存缓冲液:Preservative: 0.03% Proclin 300
Constituents: 50% Glycerol, 0.01M PBS, pH 7.4 -
产品提供形式:Liquid
-
应用范围:ELISA, GICA
-
推荐稀释比:
Application Recommended Dilution ELISA 1:10000-1:50000 GICA 1:500-1:25000 -
Protocols:
-
储存条件:Upon receipt, store at -20°C or -80°C. Avoid repeated freeze.
-
货期:Basically, we can dispatch the products out in 1-3 working days after receiving your orders. Delivery time maybe differs from different purchasing way or location, please kindly consult your local distributors for specific delivery time.
-
用途:For Research Use Only. Not for use in diagnostic or therapeutic procedures.
相关产品
靶点详情
-
功能:attaches the virion to the cell membrane by interacting with host receptor, initiating the infection. Binding to human ACE2 receptor and internalization of the virus into the endosomes of the host cell induces conformational changes in the Spike glycoprotein. Binding to host NRP1 and NRP2 via C-terminal polybasic sequence enhances virion entry into host cell. This interaction may explain virus tropism of human olfactory epithelium cells, which express high level of NRP1 and NRP2 but low level of ACE2. The stalk domain of S contains three hinges, giving the head unexpected orientat...显示更多
-
基因功能参考文献:
- Study presents crystal structure of C-terminal domain of SARS-CoV-2 (SARS-CoV-2-CTD) spike S protein in complex with human ACE2 (hACE2); hACE2-binding mode similar overall to that observed for SARS-CoV. However, details at the binding interface show that key residue substitutions in SARS-CoV-2-CTD slightly strengthen the interaction and lead to higher affinity for receptor binding than SARS-CoV receptor-binding domain. PMID: 32378705
- crystal structure of the receptor-binding domain (RBD) of the spike protein of SARS-CoV-2 bound to the cell receptor ACE2 PMID: 32365751
- crystal structure of the receptor-binding domain (RBD) of the spike protein of SARS-CoV-2 (engineered to facilitate crystallization) in complex with ACE2 PMID: 32320687
- Out of the two isolates from India compared to the isolates from Wuhan, China, one was found to harbor a mutation in its receptor-binding domain (RBD) at position 407 where, arginine was replaced by isoleucine. This mutation has been seen to change the secondary structure of the protein at that region and this can potentially alter receptor binding of the virus. PMID: 32275855
- Structural modeling of the SARS-CoV-2 spike glycoprotein show similar receptor utilization between SARS-CoV-2 and SARS-CoV, despite a relatively low amino acid similarity in the receptor binding module. Compared to SARS-CoV and all other coronaviruses in Betacoronavirus lineage B, an extended structural loop containing basic amino acids were identified at the interface of the receptor binding (S1) and fusion (S2) domains. PMID: 32245784
- crystal structure of CR3022, a neutralizing antibody from a SARS patient, in complex with the receptor-binding domain of the SARS-CoV-2 spike (S) protein to 3.1 A; study provides insight into how SARS-CoV-2 can be targeted by the humoral immune response and revealed a conserved, but cryptic epitope shared between SARS-CoV-2 and SARS-CoV PMID: 32225176
- SARS-CoV and SARS-CoV-2 spike proteins have comparable binding affinities achieved by balancing energetics and dynamics. The SARS-CoV-2-ACE2 complex contains a higher number of contacts, a larger interface area, and decreased interface residue fluctuations relative to the SARS-CoV-ACE2 complex. PMID: 32225175
- Interaction interface between cat/dog/pangolin/Chinese hamster ACE2 and SARS-CoV/SARS-CoV-2 S protein was simulated through homology modeling. Authors identified that N82 of ACE2 showed closer contact with receptor-binding domain of S protein than human ACE2. PMID: 32221306
- SARS-CoV-2 S glycoprotein harbors a furin cleavage site at the boundary between the S1/S2 subunits, which is processed during biogenesis and sets this virus apart from SARS-CoV and SARS-related CoVs; determined cryo-EM structures of the SARS-CoV-2 S ectodomain trimer. PMID: 32201080
- Study demonstrates that SARS-CoV-2 uses the SARS-CoV receptor ACE2 for entry and the serine protease TMPRSS2 for S protein priming. PMID: 32155444
- The ACE2-B0AT1 complex exists as a dimer of heterodimers. Structural alignment of the RBD-ACE2-B0AT1 ternary complex with the S protein of SARS-CoV-2 suggests that two S protein trimers can simultaneously bind to an ACE2 homodimer. PMID: 32142651
- study demonstrated SARS-CoV-2 S protein entry on 293/hACE2 cells is mainly mediated through endocytosis, and PIKfyve, TPC2 and cathepsin L are critical for virus entry; found that SARS-CoV-2 S protein could trigger syncytia in 293/hACE2 cells independent of exogenous protease; there was limited cross-neutralization activity between convalescent sera from SARS and COVID-19 patients PMID: 32132184
- study determined a 3.5-angstrom-resolution cryo-electron microscopy structure of the 2019-nCoV S trimer in the prefusion conformation; provided biophysical and structural evidence that the 2019-nCoV S protein binds angiotensin-converting enzyme 2 (ACE2) with higher affinity than does severe acute respiratory syndrome (SARS)-CoV S PMID: 32075877
收起更多
-
亚细胞定位:Virion membrane; Single-pass type I membrane protein. Host endoplasmic reticulum-Golgi intermediate compartment membrane; Single-pass type I membrane protein. Host cell membrane; Single-pass type I membrane protein.
-
蛋白家族:Betacoronaviruses spike protein family